A correction has been published for a Communications Chemistry study describing a fluorogenic, peptide-based probe designed to detect Cathepsin D in macrophages. The original paper appeared online in November 2023, and the update was issued in January 2026.
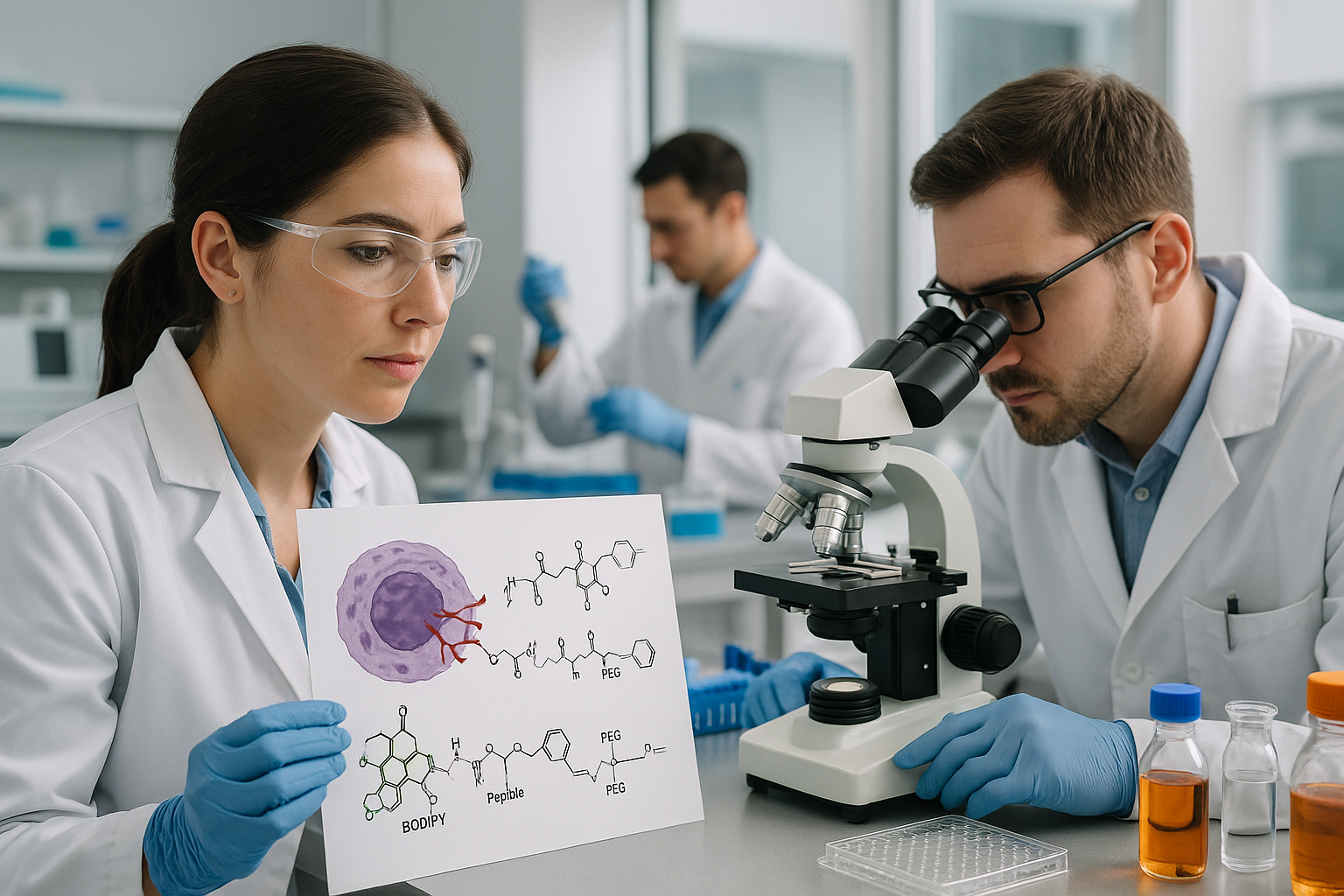
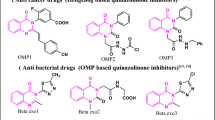
According to the notice, several structural drawings were shown incorrectly in the first version of the article. In Figure 1, the CatD-P3 probe was missing a carbon atom in the PEG tag, and the alkyne-PEG(5 K)-OMe reagent used in the click-chemistry step had been depicted with the wrong terminal group. Similar labeling and structure issues also affected Figure 2 and Supplementary Note 4.
The correction additionally states that the charges in one of the structures in Figure 2 were misplaced within the BODIPY fluorophore. Importantly, the authors and journal note that these were presentation errors only, with no effect on the experimental findings or the conclusions of the study.
The revised Figures 1 and 2, along with Supplementary Note 4, have been updated in the HTML and PDF versions of the article. The paper remains open access under a Creative Commons Attribution 4.0 International License.