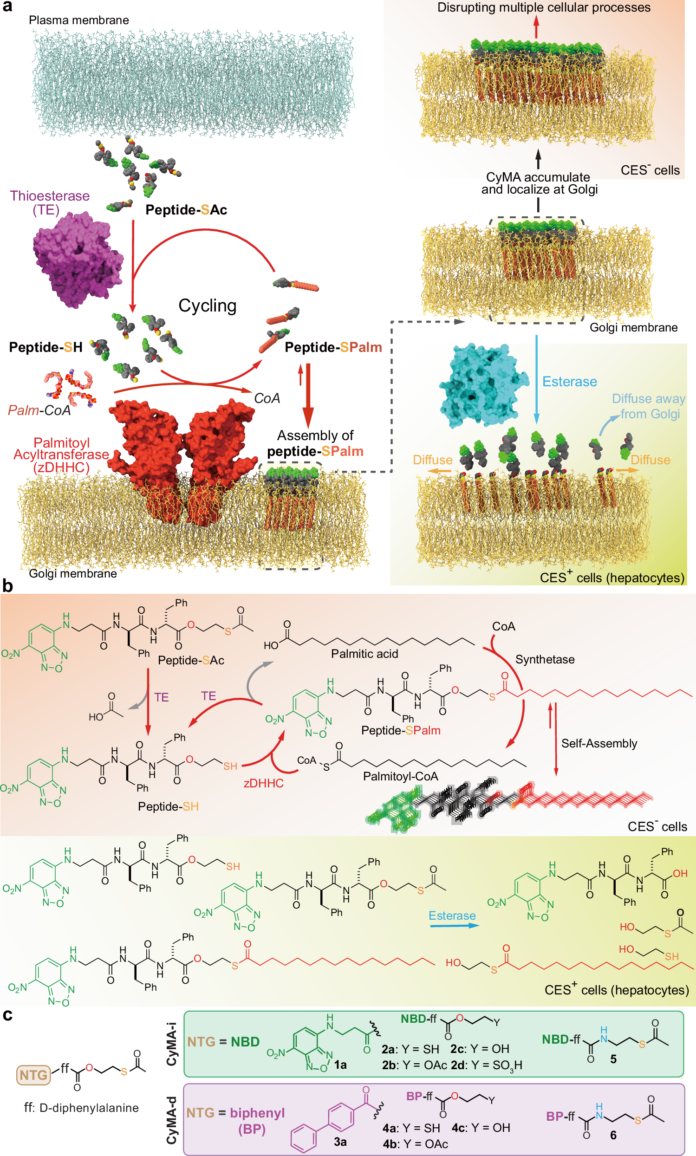

The Golgi apparatus sits at the center of protein trafficking, lipid processing, and signaling, making it an appealing but difficult target for both imaging and therapeutic intervention. A new peptide-based strategy called cycling molecular assemblies, or CyMA, uses a built-in enzymatic loop to localize at the Golgi and either visualize it quickly or interfere with its function.
The approach starts with amphiphilic peptide thioesters that can enter cells. Once inside, thioesterases remove the protecting group to expose a thiol, which is then re-acylated by Golgi-resident palmitoyltransferases. This back-and-forth chemistry creates a dynamic, Golgi-enriched assembly that is effectively trapped in place by continuous cycling through deacylation and palmitoylation.
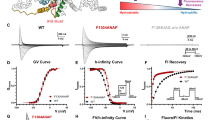
For imaging, the CyMA design produces near-immediate Golgi signal, even at very low probe concentrations. In a second version, the fluorescent component is replaced with a biphenyl group, converting the system from a reporter into a functional disruptor. This non-fluorescent variant accumulates at the Golgi and impairs protein modification, trafficking, and secretion, ultimately reducing cell proliferation and causing cell death in experimental models.
One notable feature of the platform is selectivity: the disruptor appears to spare some cell types, including hepatocytes and certain immune cells, likely because of differences in esterase activity. That makes the work interesting not only as a Golgi-targeting method, but also as a proof of concept for using enzyme-driven supramolecular assemblies to selectively trap peptide materials at specific organelles.
More broadly, the study suggests a modular framework that could be adapted to other cellular compartments by swapping in different enzyme pairs. In other words, dynamic peptide assembly may become a general way to image, perturb, or exploit organelles through chemistry that is actively maintained inside living cells.