Targeted protein degradation has reshaped drug discovery by making it possible to eliminate proteins rather than merely block them. Most of the best-known degraders, including PROTACs, work by bringing a target protein into close contact with an endogenous E3 ubiquitin ligase, which then tags the target for destruction by the proteasome.
That approach has proven powerful, but it also comes with design challenges. The target, linker, and E3 ligase must all fit together in a productive geometry for ubiquitination to occur efficiently. In practice, that can make optimization slow and unpredictable. It can also create a route to resistance if the ligase loses activity or if mutations disrupt the target–ligase interface.
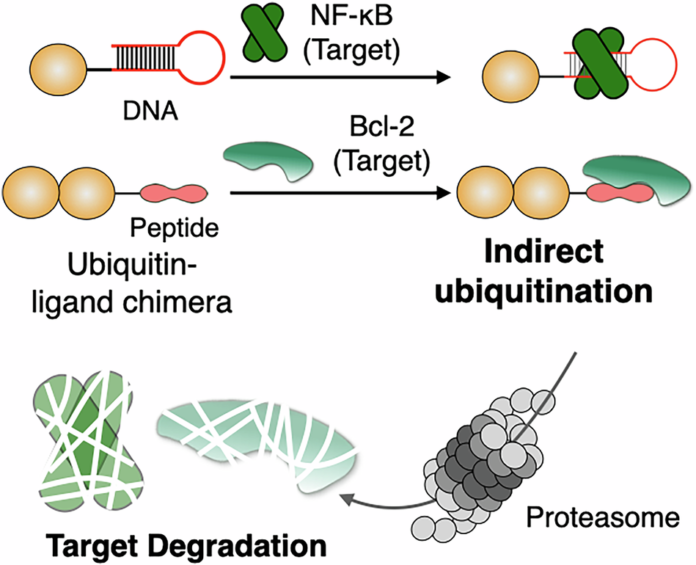
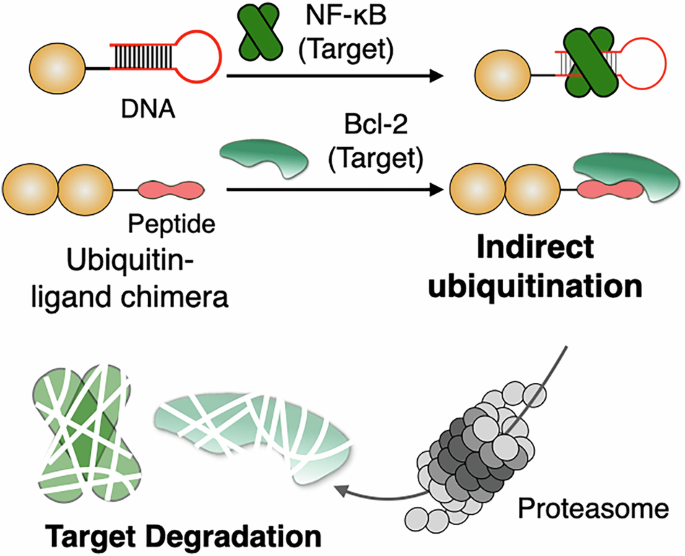
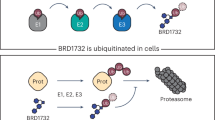
In a new study, researchers report an alternative they call indirect ubiquitination. Instead of depending on the cell’s own ubiquitination machinery to build the degradation signal, the strategy uses a chimeric molecule made from a target-binding ligand and a ubiquitin moiety. Through non-covalent binding to the target protein, the system places ubiquitin in position to promote degradation without requiring a recruited endogenous E3 ligase.
The team showed that this design could drive proteasomal breakdown of recombinant Bcl-2 and NF-κB p50, and they also demonstrated degradation of endogenous Bcl-2 inside cells. Those results suggest that ubiquitin itself can be used more directly as a chemical handle for protein removal, opening a different path for designing degraders.
The concept could matter for targets that are difficult to address with conventional PROTACs, especially when E3 availability, ligase expression, or ternary complex formation limits performance. If further developed, indirect ubiquitination may expand the toolkit for targeted protein degradation with a platform that is less dependent on the cell’s native E3 landscape.
For peptide and protein engineering researchers, the work is notable because it reframes ubiquitin not just as a downstream signal, but as a modular component that can be chemically delivered to a target protein. That may create new opportunities for designing next-generation proteolysis tools with different binding requirements and resistance profiles.