Solid-phase peptide synthesis has long been the workhorse for making peptides used in drug discovery, biomaterials research, and applications in agriculture, veterinary medicine, and cosmetics. But the method has a sustainability problem: it typically relies on large amounts of hazardous organic solvents and disposable solid supports that add to the waste stream.
That challenge has become more urgent as solvent-reduction rules tighten and labs look for practical replacements for conventional peptide chemistry. The newest work highlighted in Nature Sustainability points to a promising path forward: moving more of SPPS into water.
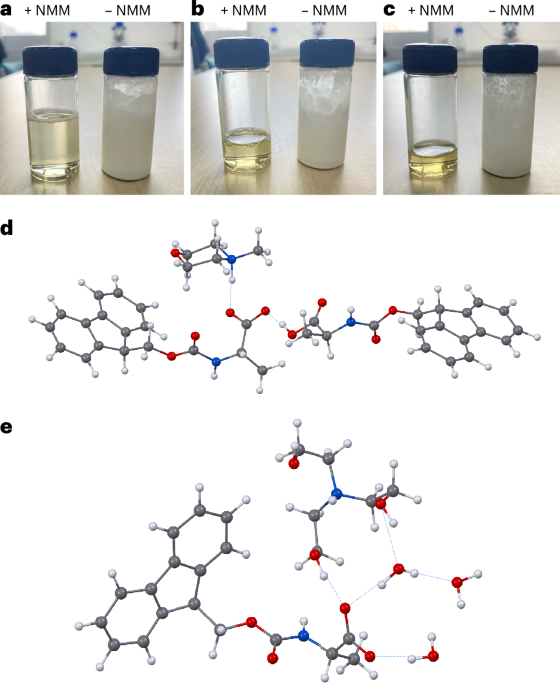
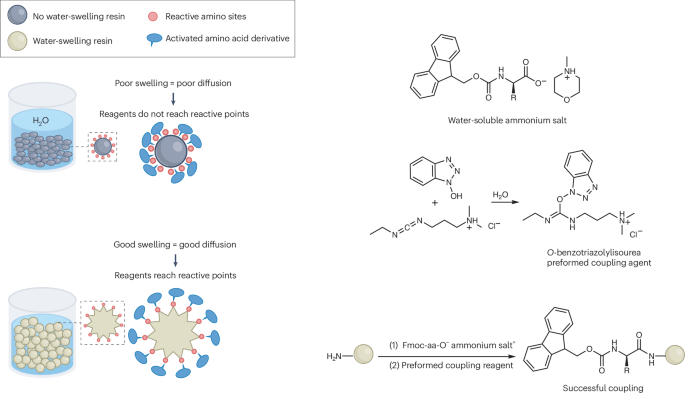
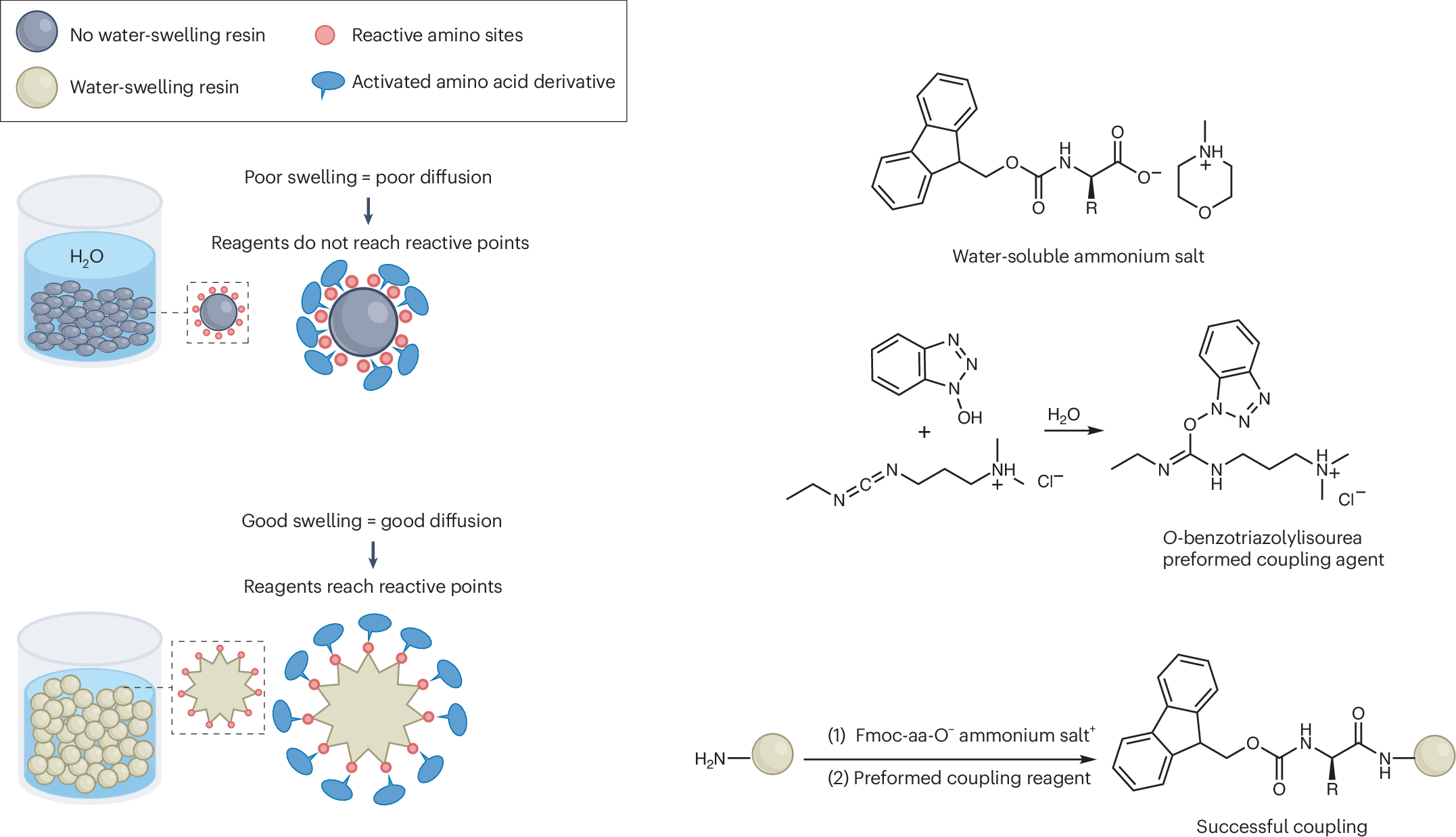
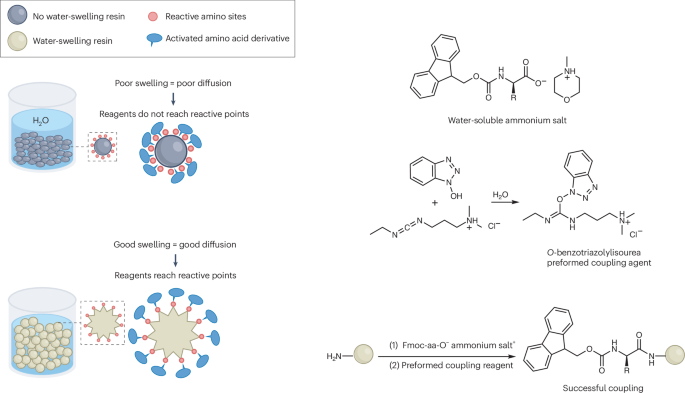
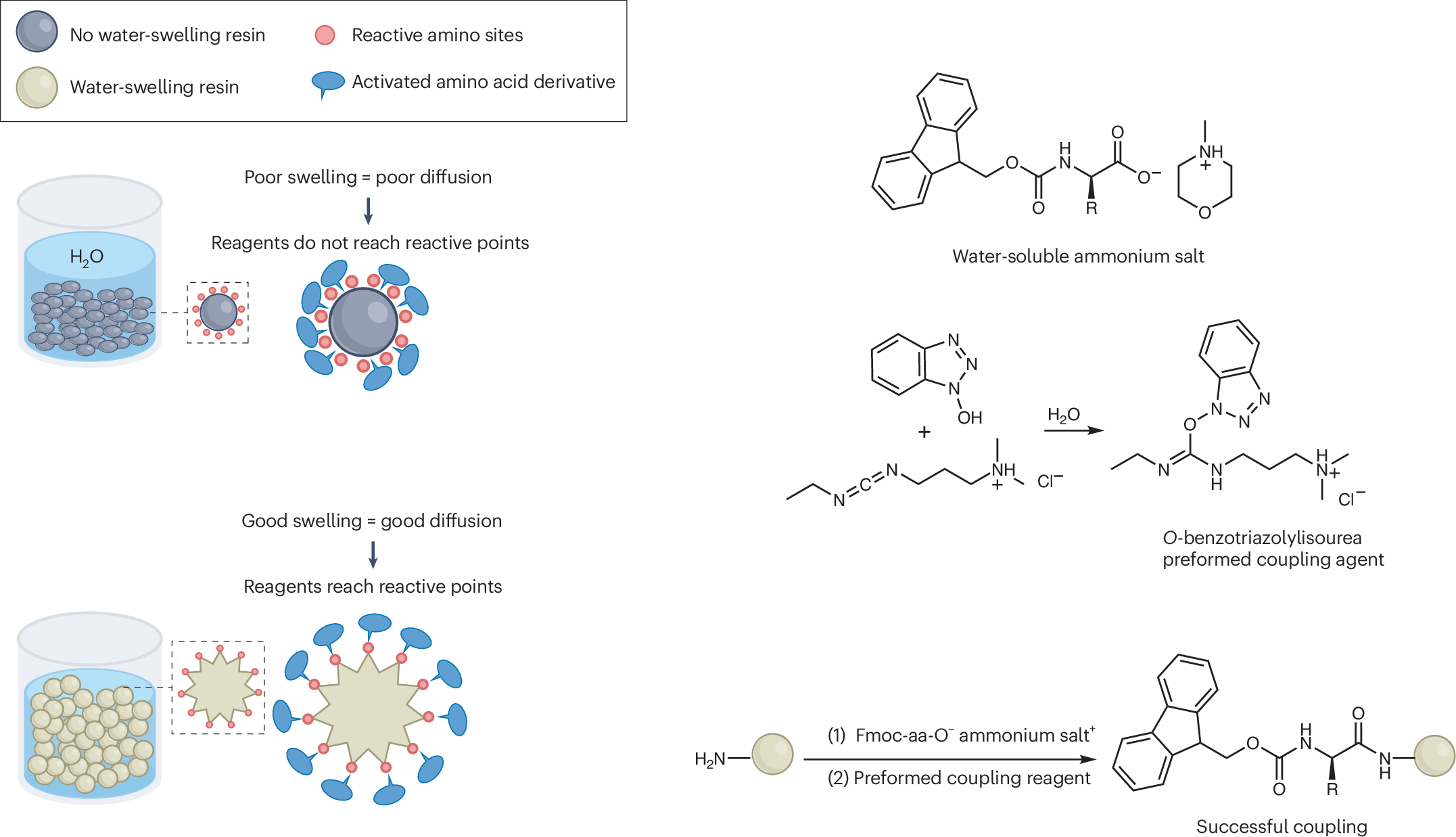
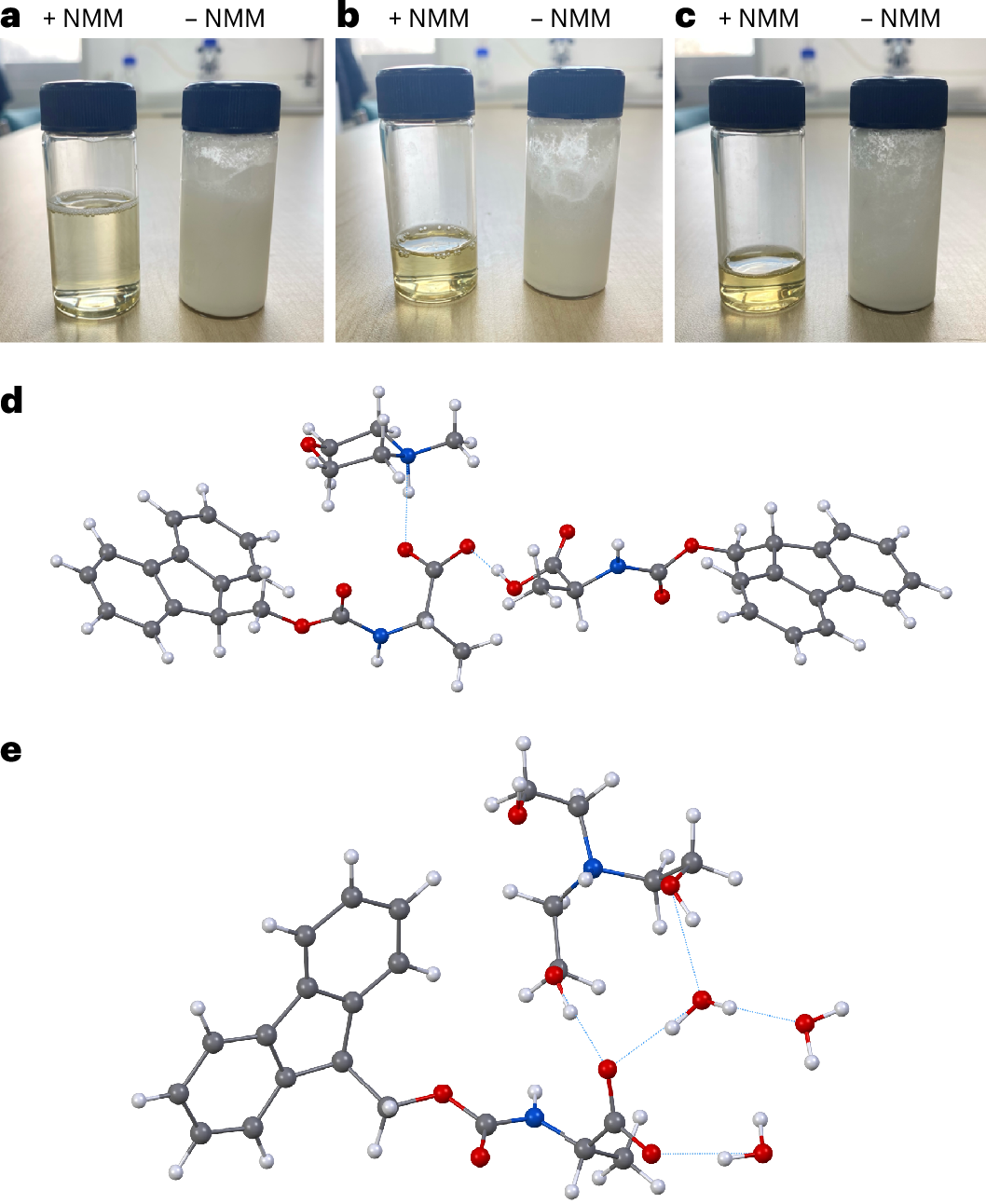
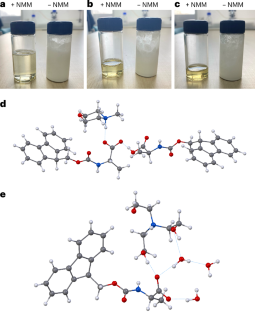
The key obstacle has always been the starting materials. The standard Nα-Fmoc-protected amino acids used in SPPS do not dissolve well in water, which makes an aqueous process difficult to run at useful concentrations. In this study, the researchers found that pairing these protected amino acids with certain amine salts, such as those formed from N-methylmorpholine or triethanolamine, produces highly water-soluble mixtures.
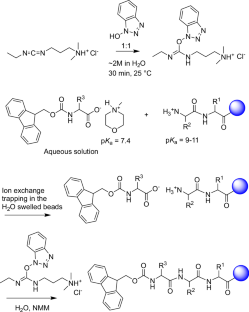
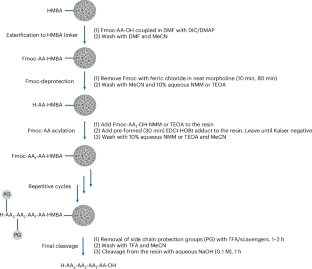
With that solubility issue addressed, the team combined the amino-acid feedstock with a pre-formed water-soluble activating agent and carried out peptide assembly on a hydrophilic, biodegradable poly-ε-lysine-based solid support. According to the report, the reaction proceeds cleanly in aqueous solution and avoids the side reactions that often complicate peptide synthesis.
Why this matters: if the approach scales, it could reduce dependence on toxic solvent systems and non-degradable resins without sacrificing the practicality that made SPPS so widely adopted in the first place. For peptide research, that would mean a more sustainable way to build the molecules that underpin many modern therapeutics and biological studies.
The broader significance is simple but important. Peptide manufacturing has been searching for greener chemistry that still works with real-world synthesis demands. Water-based SPPS will not replace every existing workflow overnight, but this study shows that the gap between performance and sustainability may be narrowing.