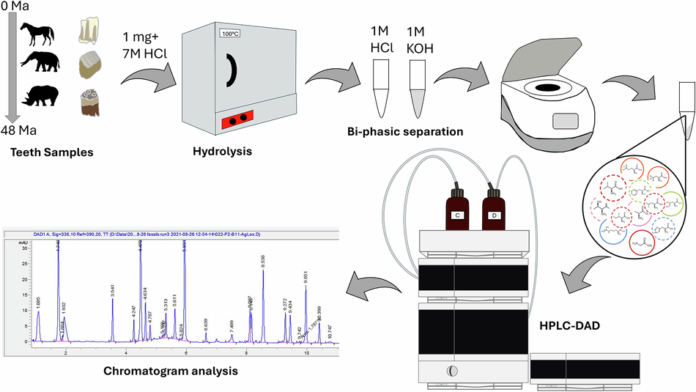
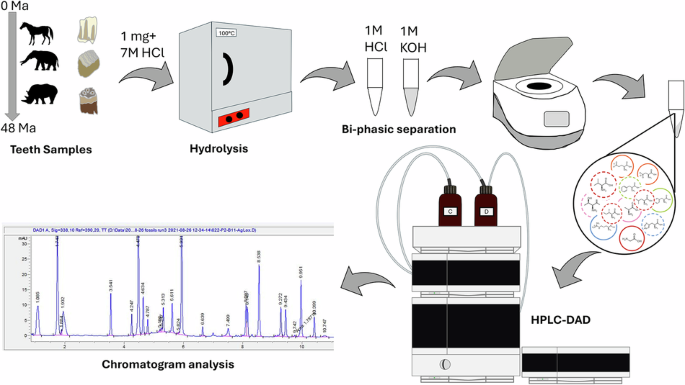
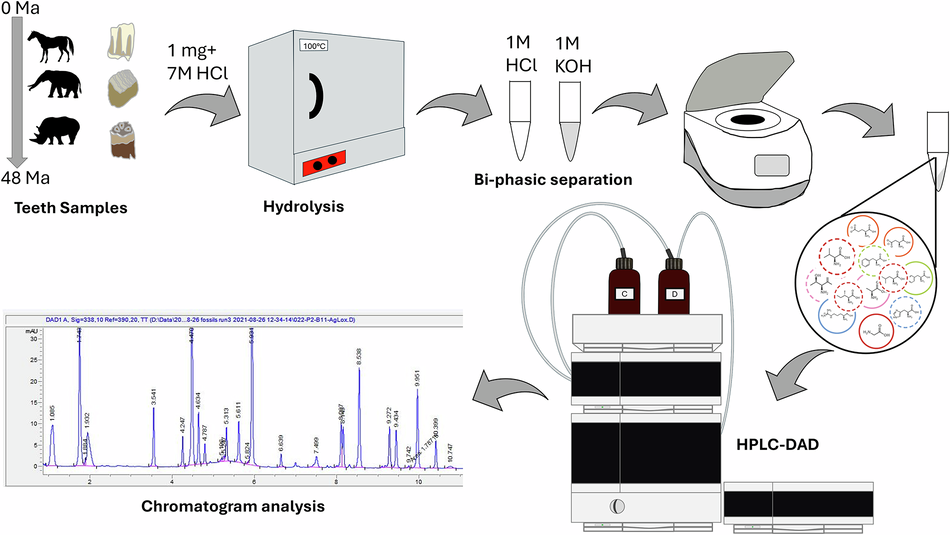
Tooth enamel has long been viewed as one of the most durable tissues in the fossil record, but a new study pushes that reputation much further. Researchers examining modern and fossil enamel from large herbivorous mammals found that total hydrolysable amino acids can persist in enamel for at least 48 million years, extending well into the Eocene.
The work focused on enamel from horses, rhinos, and proboscideans across a range of fossil ages and preservation settings. The key takeaway is that enamel does not simply retain a faint chemical shadow of past biology: it can keep a measurable endogenous amino-acid signal over deep time.
A major pattern in the study is an early drop in the organic signal, followed by long-term stabilization of what the authors describe as the intra-crystalline fraction. In other words, once the most exposed material is lost during the earliest phase of burial and alteration, the amino acids trapped within enamel crystals appear to behave like a closed system.
That matters because it suggests enamel is far more robust than many other fossil tissues when it comes to preserving organic information. The study also reports that preservation was not strongly tied to taphonomic context, meaning the amino-acid patterns were similar across different burial and environmental histories. Relative amino-acid abundances were variable in both modern and fossil samples, but the overall persistence of the signal remained clear.
For paleontology and biomolecular research, the implications are significant. If enamel can retain endogenous amino acids this deeply in time, it could become an even more valuable archive for reconstructing evolutionary relationships, diet, and ancient ecosystems. The findings also strengthen interest in enamel as a source of durable biomolecular evidence, complementing earlier work showing that fossil teeth can preserve peptides and nitrogen-isotope signals.
In short, mammalian tooth enamel may be one of the most reliable natural vaults for ancient organic matter, preserving amino-acid traces long after most other biomolecules have disappeared.