Designing peptide substrates for proteases has always been a difficult mix of biology, chemistry, and brute-force screening. The challenge is not just finding a sequence a protease can cut, but finding one that is cut efficiently and selectively. With an enormous sequence space and limited experimental throughput, substrate discovery has often relied on literature mining, expert intuition, or expensive large-scale screens.
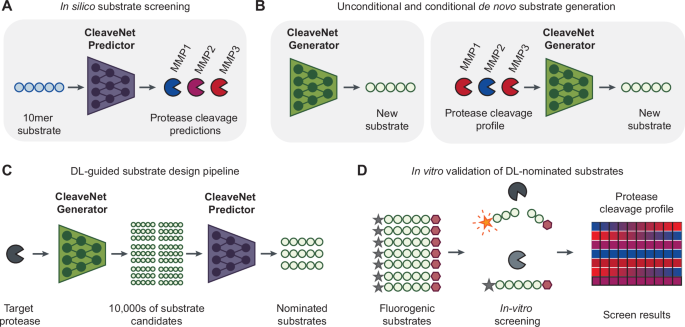
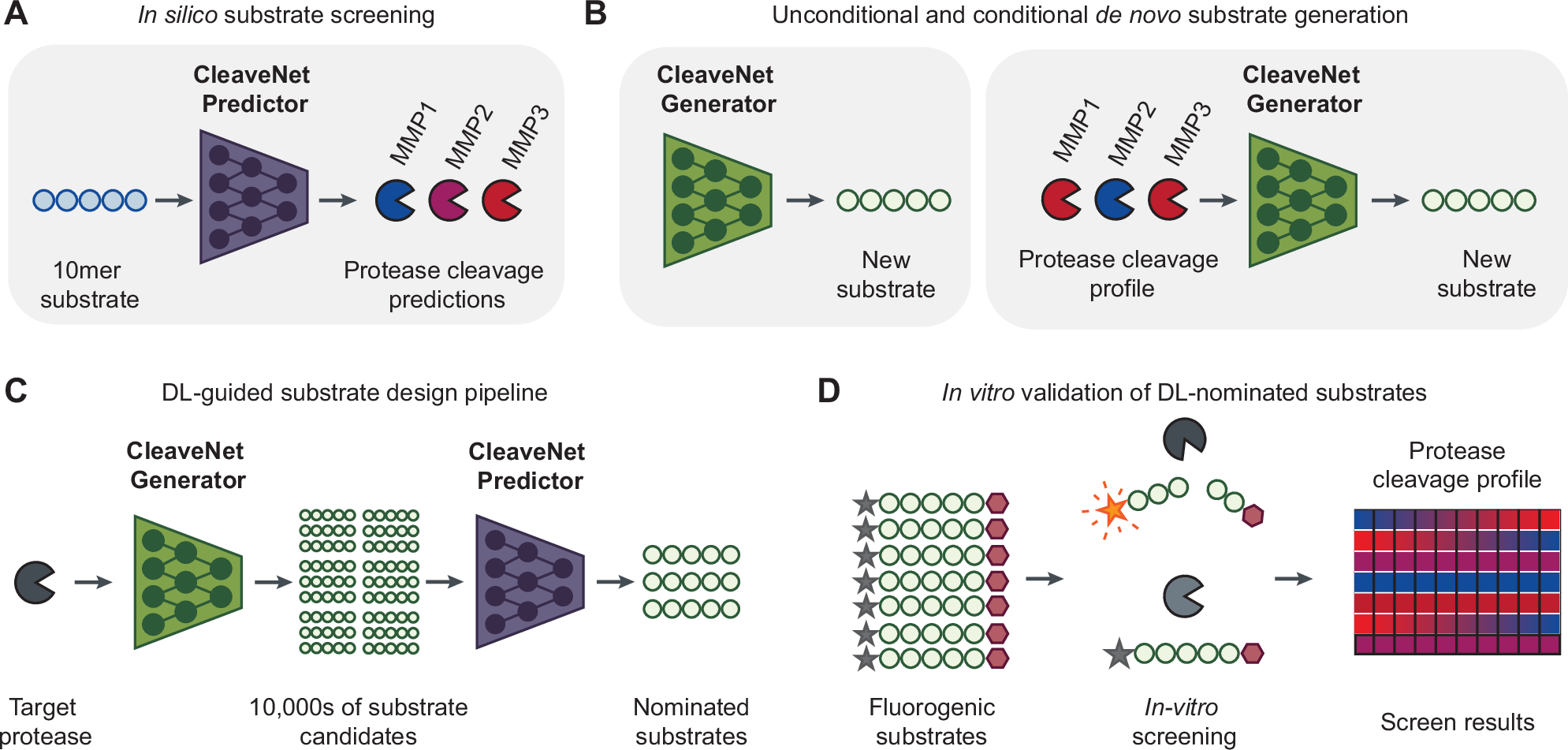
A new study introduces CleaveNet, an end-to-end deep learning pipeline built to address that bottleneck. Rather than only predicting whether a peptide will be cleaved, the system is designed to generate candidate substrates directly. In other words, it aims to move from passive prediction to active design.
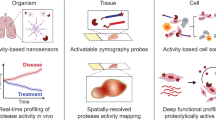
The method was applied to matrix metalloproteinases, a family of proteases with important roles in tissue remodeling, inflammation, and cancer. According to the study, CleaveNet can produce peptide substrates with favorable biophysical properties while also capturing known cleavage preferences and uncovering motifs that had not been well characterized before.
One notable feature is a conditioning tag that allows users to steer generation toward desired cleavage behavior. That means the model can be guided to favor more efficient substrates or, importantly, more selective ones. Selectivity matters because closely related proteases often recognize overlapping peptide patterns, making it hard to design probes that respond to just one target.
The researchers validated the generated peptides in a large in vitro screen, including the especially demanding case of designing selective substrates for MMP13. Experimental results supported the idea that AI-assisted design can reduce the trial-and-error burden normally associated with protease substrate development.
For peptide researchers, the broader significance is clear: if models like CleaveNet continue to improve, they could shorten the path from enzyme biology to usable probes, diagnostic triggers, and therapeutic tools. The approach may also point toward a more general future for enzyme engineering, where deep learning helps explore chemical space that is simply too large to search by hand.
In a field where substrate design has long been constrained by scale and specificity, this work suggests that generative AI may become a practical discovery engine rather than just an analytical aid.