Host defense peptides have long been viewed as promising antibiotic alternatives, especially against pathogens that have already learned to outmaneuver many small-molecule drugs. But these peptides come with a familiar problem: they are often fragile in the body and can be rapidly broken down by proteases.
A common fix is retro-inversion, a design strategy that flips both the peptide backbone direction and the amino acid chirality. The goal is to preserve the overall side-chain arrangement while making the molecule harder to degrade. For many bacteria, however, this approach has been a mixed bag, because the structural changes can also reduce biological activity.
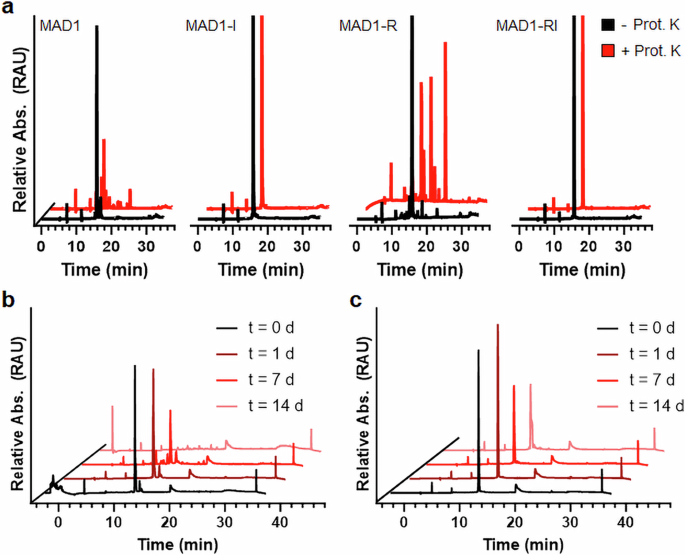
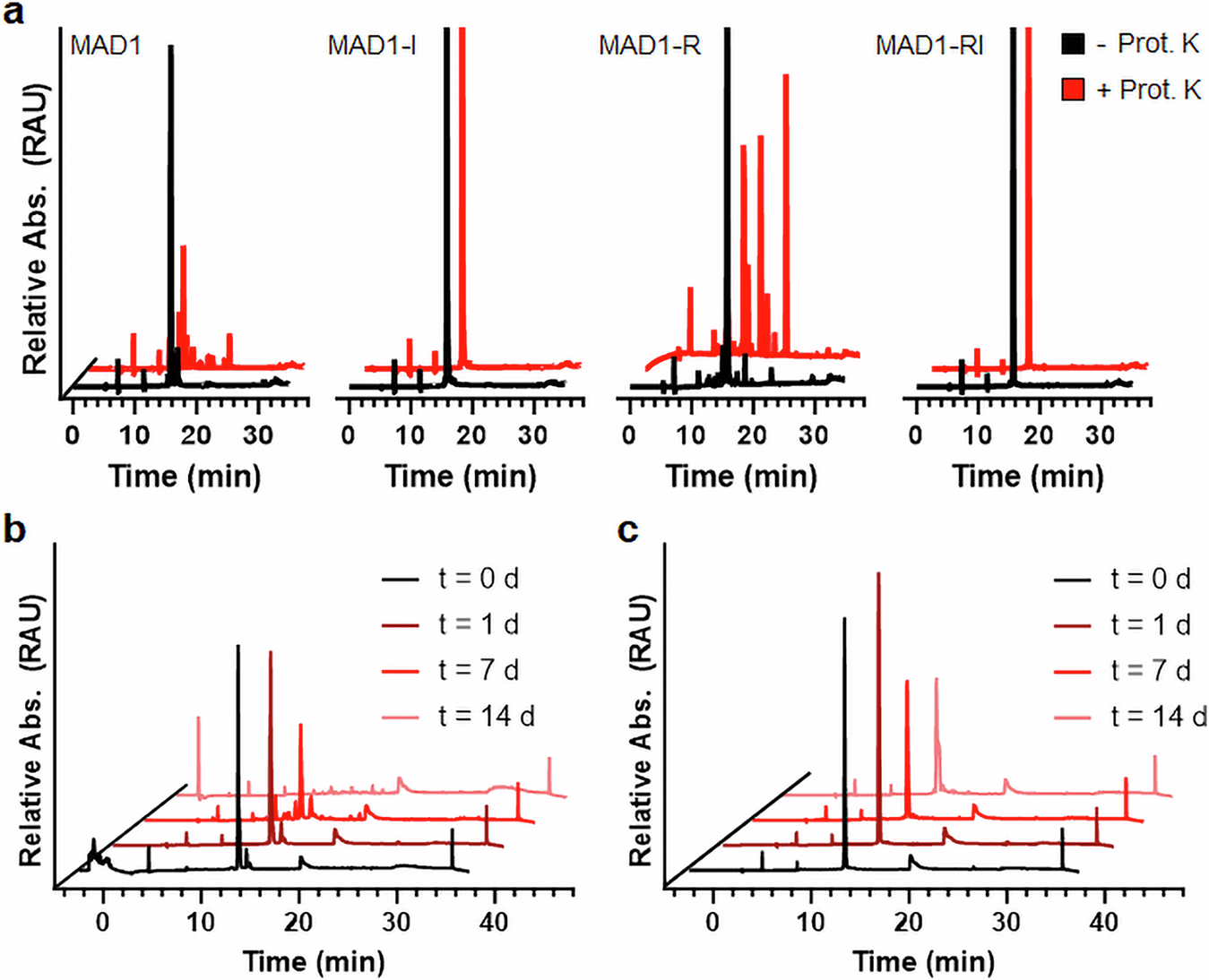
This new study turns that assumption on its head for mycobacteria. The researchers found that retro-inverted versions of antimycobacterial host defense peptides were not merely more stable, but in many cases substantially more potent, more selective, and safer for host cells. In some comparisons, activity improved by more than tenfold.
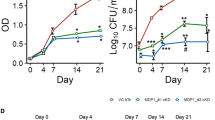
The standout candidate, called MAD1-RI, showed especially strong performance. It rapidly sterilized replicating Mycobacterium tuberculosis cultures, worked against drug-resistant clinical isolates, and boosted the effects of co-administered tuberculosis antibiotics.
Mechanistic experiments pointed to an unexpected explanation. The gains in activity did not appear to come mainly from improved resistance to proteolysis. Instead, the retro-inverted peptides seemed to alter the physical behavior of the mycobacterial membrane, changing membrane thermodynamics in a way that made the cells more vulnerable.
Transcriptomic analyses also suggested that MAD1-RI attacks mycobacteria through a combination of membrane disruption and metabolic stress. Taken together, the findings suggest that retro-inversion may be more than a stability trick: for antimycobacterial peptide design, it could be a powerful way to unlock selectivity and potency.
Because tuberculosis and non-tuberculous mycobacterial infections continue to rise, including in drug-resistant forms, the study adds a timely tool to the peptide antibiotic toolkit. It also highlights an important reminder for peptide engineering: a modification that weakens activity in one pathogen may enhance it in another.