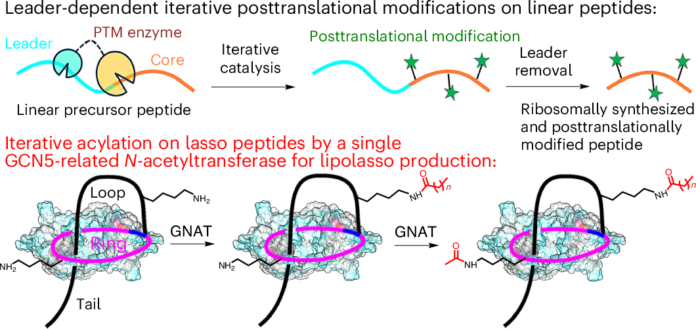
Ribosomally synthesized and post-translationally modified peptides, or RiPPs, are prized for the chemical variety they can achieve from relatively small genetic blueprints. A new study shows that this diversity can be expanded even further after a peptide has already folded into one of the most architecturally constrained RiPP classes: the lasso peptide.
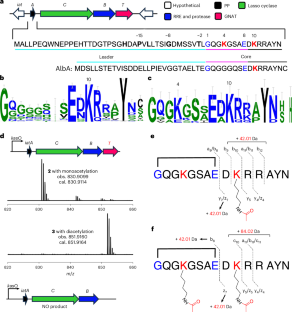
In the work, the authors describe a widespread group of GCN5-related N-acetyltransferases, or GNATs, that do something unusual for iterative enzymes. Instead of acting on a flexible linear peptide, these enzymes can recognize a mature lasso peptide and acetylate two lysine residues one after the other, placing modifications on both the loop and ring regions of the folded molecule.
That matters because lasso peptides are not easy substrates. Their threaded, knotted-like topology creates a compact three-dimensional shape that typically demands highly tailored enzyme-substrate interactions. To understand how one of these enzymes, IatT, handles such a challenging target, the researchers combined enzymatic reconstitution with high-resolution cryo-electron microscopy.
The structural work revealed the binding pocket used by IatT to engage the lasso peptide and highlighted residues that help separate the first acetylation event from the second. In other words, the enzyme appears to use a single active site architecture to carry out two related but distinct steps on the same folded substrate.
The most practical result came from protein engineering. By reshaping the acetyl-recognition site, the team expanded the cavity enough to accept longer acyl groups. That turned the enzyme into a tool for generating new acylated lasso peptides, including a newly framed category the authors describe as lipolasso peptides.
Because GNAT-encoding genes are common across RiPP biosynthetic gene clusters, the strategy could have broad reach. Rather than discovering one-off pathways for each new scaffold, researchers may be able to repurpose similar enzymes to diversify peptide natural products more efficiently and create ribosomal lipopeptides with new properties.
The study adds to a growing picture of RiPP biosynthesis as a modular platform for late-stage editing. It also shows that mature, highly structured peptides are not necessarily off-limits to iterative tailoring enzymes, opening a route to engineered products that sit at the intersection of peptide natural products and lipopeptide chemistry.