Histone acetyltransferases help control gene expression by loosening chromatin and making DNA more accessible to the transcriptional machinery. But these enzymes rarely work in isolation. Instead, they act within multi-protein complexes that shape where they go in the genome, which substrates they modify, and how they influence cell behavior.
That complexity creates a challenge for drug discovery. If a compound blocks the shared catalytic enzyme, it may hit several related complexes at once, making it difficult to tease apart which biological effects come from which assembly. In a new study, researchers took a more selective approach: rather than targeting the common HAT enzyme, they designed inhibitors against YEATS2, a subunit unique to the ATAC complex.
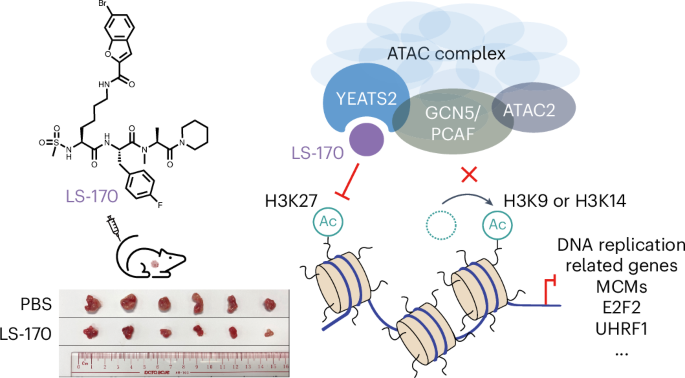
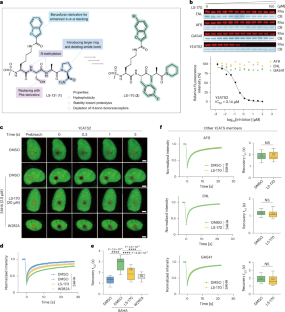
The goal was to disrupt ATAC specifically and observe what happens when this complex is removed from chromatin. The lead compound, LS-170, showed the strongest activity. According to the study, it reduced ATAC occupancy on chromatin, lowered ATAC-dependent histone acetylation, and suppressed expression of genes controlled by ATAC.
Those molecular changes also translated into a functional effect in vivo. In a lung cancer mouse model, treatment with the inhibitor significantly restrained tumor growth, suggesting that ATAC may be a promising therapeutic vulnerability in at least some cancer settings.
Beyond the therapeutic angle, the work provides a useful chemical tool for studying how HAT complexes behave in cells. Because ATAC is part of a broader family of acetyltransferase assemblies with overlapping enzymatic features, a complex-selective inhibitor could help researchers map out which genes and pathways depend on ATAC specifically, rather than on HAT activity in general.
The study also adds to a growing view in chromatin biology: the composition of a protein complex can be just as important as the enzyme it carries. By targeting a complex-specific subunit, the authors demonstrate a way to dissect chromatin regulation with greater precision and potentially open a new route for epigenetic drug development.
Structural data associated with the work were deposited in the Protein Data Bank, and sequencing datasets were made publicly available through GEO.